Citation Discovery Tools for Conducting Adaptive Meta-analyses to Update Systematic Reviews
Article information
Abstract
Objectives:
The systematic review (SR) is a research methodology that aims to synthesize related evidence. Updating previously conducted SRs is necessary when new evidence has been produced, but no consensus has yet emerged on the appropriate update methodology. The authors have developed a new SR update method called ‘adaptive meta-analysis’ (AMA) using the ‘cited by’, ‘similar articles’, and ‘related articles’ citation discovery tools in the PubMed and Scopus databases. This study evaluates the usefulness of these citation discovery tools for updating SRs.
Methods:
Lists were constructed by applying the citation discovery tools in the two databases to the articles analyzed by a published SR. The degree of overlap between the lists and distribution of excluded results were evaluated.
Results:
The articles ultimately selected for the SR update meta-analysis were found in the lists obtained from the ‘cited by’ and ‘similar’ tools in PubMed. Most of the selected articles appeared in both the ‘cited by’ lists in Scopus and PubMed. The Scopus ‘related’ tool did not identify the appropriate articles.
Conclusions:
The AMA, which involves using both citation discovery tools in PubMed, and optionally, the ‘related’ tool in Scopus, was found to be useful for updating an SR.
INTRODUCTION
A systematic review (SR) is a research methodology that synthesizes the findings of studies with the same research hypothesis to produce a summary conclusion on the hypothesis [1]. The SR process involves a comprehensive search of related evidence, followed by a meta-analysis, which summarizes the extracted results. The literature search for an SR is conducted through an electronic search, which applies a search algorithm comprised of related keywords related by Boolean logic (AND, OR, NOT) in databases such as PubMed (www.ncbi.nlm.nih.gov/pubmed/) [2]. The publication date of research articles is reflected as a limitation in search conditions because of depending on the timing of the literature search. Furthermore, updates should be conducted regularly to reflect new evidence that has been published after the literature search was conducted for the previous published SR [3,4].
In the 2008 Cochrane Handbook for Systematic Reviews of Interventions, no specific updating methodology is mentioned, although the handbook requires updates to be conducted within two years of the initial publication of the SR [5]. However, this recommendation is rarely followed [6,7]. This is because methodologies for performing the update have not yet been established [8,9], and the update requires human resources and time [4]. For topics in which the number of searching engines is increasing, the burden of updating necessarily increases as well [10]. Therefore, methodologies that enable more efficient searches are needed.
PubMed and Scopus (www.scopus.com), which are the main databases searched for SRs, each have citation discovery tools that provide a list of articles that cited the article of interest and a list of similar or related articles. We developed an SR update methodology that uses these tools. Lists of articles citing or related to the existing SR are first constructed from the citation discovery tools without a separate electronic search, and then articles to be used in the new meta-analysis are selected [11-14]. This methodology assumes that studies conducted with the same research hypothesis have a high likelihood of citing the articles included in the original SR or of being in the list of similar/related articles, and thus that our method will produce a similar list to what the Cochrane review update (CRU) method would produce. In order to differentiate the new methodology from the CRU, which reproduces the SR according to a previously developed protocol, we have named this methodology ‘adaptive meta-analysis’ (AMA) [11-14]. The AMA approach to SR updating was developed on the basis of the techniques for using the citation discovery tools we found to be most appropriate.
The ‘related articles’ citation discovery tool in Scopus produces lists of more than 5000 citations per article, which makes the search too broad to be useful. On the other hand, it is unclear whether the more targeted citation discovery tools that produce shorter lists are inclusive enough. It is worth determining whether using the citation discovery tools of Scopus and PubMed without performing other searches is sufficient to produce a comprehensive list of the articles that should ultimately be included in an SR update. Thus, this study aims to evaluate the value of using the citation discovery tools following the AMA method for an SR update.
METHODS
In order to investigate the usefulness of the ‘cited by’ citation discovery tool of Scopus, we selected the SR published by one of the present co-authors [15] in 2008 because the SR needed an update and the SR authors had already published a summary of findings published later on the hypothesis covered by the SR [12]. The electronic databases used by Bae et al. [15] for the original SR were PubMed, Ovid MEDLINE, and Embase. The search terms used were ([pancreas] OR [pancreatic]) AND ([neoplasm] OR [cancer]) AND ([fruit] OR [citrus]). For each of 10 articles (nine articles and the SR of Bae et al. [15]), we used the citation discovery tools to produce lists of ‘cited by’ articles provided by PubMed (PM_C), ‘similar articles’ provided by PubMed (PM_S), and ‘cited by’ articles provided by Scopus (SC_C). The long lists of ‘related articles’ provided by Scopus (SC_R) were used just to confirm the number of articles included in the lists. The latest publication date included in the search was late November 2015.
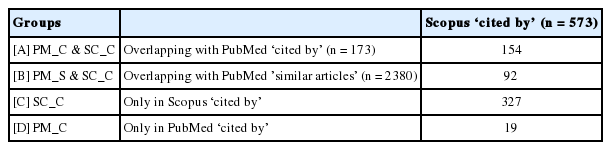
The three types of lists were divided into the following four groups: (A) lists of articles in both PM_C and SC_C, (B) lists of articles in both PM_S and SC_C, (C) lists of articles included only in SC_C, and (D) lists of articles included only in PM_C.
The inclusion criteria for the updated meta-analysis were as follows: analytical epidemiological studies (i.e., case-control studies or cohort studies) that investigated whether pancreatic cancer risk decreased with the level of citrus fruit intake. We evaluated the abstracts and full texts of articles included in the three lists (PM_C, PM_S, SC_C), and then excluded articles according to the following criteria: (1) laboratory studies on animal, cellular, and genetic levels, (2) studies on diseases other than pancreatic cancer, (3) studies that involved developing research methodologies, (4) studies on disease statistics, (5) expert opinions or reviews, (6) systematic reviews, (7) observational studies without information on citrus fruit intake, and (8) clinical studies of patients. According to the aforementioned criteria, we investigated the distribution of articles by the three citation discovery tool lists. The distribution of the selected articles was used as an index for evaluation of the usefulness of the citation discovery tools for the SR update.
RESULTS
When the ‘cited by’, ‘similar articles’, and ‘related articles’ citation discovery tools in PubMed and Scopus were applied to the nine articles selected in the meta-analysis by Bae et al. [15] plus the article by Bae et al. itself, the PM_C lists contained a total of 173 articles, while the PM_S lists included 2380 articles. The SC_C lists contained 573 articles, while the SC_S lists comprised 95 365 articles. The fact that there were approximately 9500 articles in the SC_S list for each article confirmed that the Scopus ‘related articles’ tool produced a search that was too broad to be useful.
Table 1 shows the overlap between the 573 articles in the SC_C lists and the PM_C and PM_S lists. Among the 173 articles in PM_C, 154 (89.1%) articles overlapped with SC_C; among the 2380 articles in PM_S, 92 (3.9%) articles overlapped with SC_C. In other words, among the 573 articles in SC_C, 246 (= 154 + 92) articles were also included in PM_C and PM_S, showing an overlap of 42.9% (= 246 / 573).
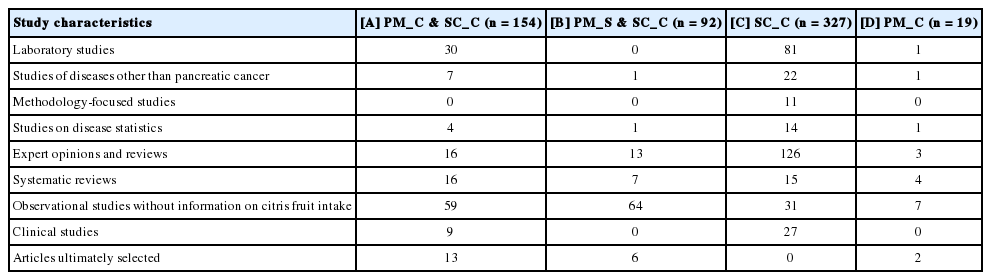
Table 2 shows the distribution of the articles according to their characteristics after the aforementioned exclusion criteria were applied. Among the four groups shown in Table 1, the group that contained the highest number of ultimately selected articles was the set of overlapping articles in PM_C and SC_C. In contrast, in the group of articles included only in SC_C, no article was selected for the meta-analysis update.
DISCUSSION
The finding that no article was selected from the list of articles included only in SC_C, which is also shown in Table 2, signifies that the ‘cited by’ and ‘similar articles’ tools provided by PubMed are sufficient for SR updates. However, since many articles were selected from the list of articles that appeared in both PM_C and SC_C, the efficiency of searches can be increased by using the overlapping articles from the two lists. We recommend this approach as the search strategy for the SR update method we have named the ’AMA’ [11-14].
For CRU, confirmation of the necessity of updates, determination of the timing of updates, and cumulative meta-analysis have been discussed [3,16]. Moreover, additional complications, such as the need to reflect changes in the calculation of risk of bias due to updates, have been suggested [4]. Given that consensus has not yet been reached on the proposed CRU update methodology, we present the AMA as a more feasible alternative to the CRU (Table 3). A CRU involves performing literature searches repeatedly at fixed intervals, using the search algorithms from the previous meta-analysis protocol [5]. In contrast, the AMA method develops a list of articles to analyze based on the articles that were selected in the original SR, which had involved searches using a stated algorithm. AMA compiles an updated list of articles for analysis by producing lists of articles that cite or are similar to the articles included in the SR, using citation discovery tools. Therefore, the most notable difference between the two methods is that CRU values protocols to generate reproducible results while AMA values lists of selected articles to generate acceptable results. Since updating an SR can mean that a meta-analysis must be conducted again on additionally selected articles that are consistent with a certain research hypothesis, we used the term ‘adaptive’ to distinguish our method from simply ‘updating’ the SR.

Differences between the Cochrane review update (CRU) and adaptive meta-analysis (AMA) for updating a systematic review (SR)
AMA has several advantages. First, omitting electronic searches reduces the time and effort needed to update an SR. The fact that all articles that need to be selected can be found using only the two citation discovery tools of PubMed is encouraging. Second, AMA is also useful for identifying included articles that report on the same cases or cohort [12]. When the ‘cited by’ and ‘similar articles’ tools are used, duplicates among published articles can be easily assessed so that the list of articles to be including in the meta-analysis can be modified as needed. Third, since the AMA method can confirm errors in previously published SRs and can additionally conduct a sensitivity analysis or subgroup analysis while adding articles that were omitted in the previous SRs, it can be used as a methodology for meta-epidemiology [17].
The validity of AMA remains to be investigated. AMA cannot yet be directly compared to other methods since there is no consensus on the criteria for including articles in a meta-analysis. Nevertheless, when AMA was applied to previous SRs in which a hand search was conducted after an electronic search, we confirmed that several articles that should have been selected, when considering the search period, were omitted [11,13,14]. This indicates that AMA can increase the accuracy in a SR literature search. In the future, it is expected that applying AMA to more studies will show that AMA is useful as a methodology for updating SRs.
Notes
CONFLICT OF INTEREST
The authors have no conflicts of interest associated with the material presented in this paper.