Articles
- Page Path
- HOME > J Prev Med Public Health > Volume 54(5); 2021 > Article
-
Systematic Review
A Systematic Review of Spatial and Spatio-temporal Analyses in Public Health Research in Korea -
Han Geul Byun*
 , Naae Lee*
, Naae Lee* , Seung-sik Hwang
, Seung-sik Hwang
-
Journal of Preventive Medicine and Public Health 2021;54(5):301-308.
DOI: https://doi.org/10.3961/jpmph.21.160
Published online: August 26, 2021
Department of Public Health Sciences, Graduate School of Public Health, Seoul National University, Seoul, Korea
- Corresponding author: Seung-sik Hwang Department of Public Health Sciences, Graduate School of Public Health, Seoul National University, 1 Gwanak-ro, Gwanak-gu, Seoul 08826, Korea E-mail: cyberdoc@snu.ac.kr
- *Byun & Lee contributed equally to this work as joint first authors.
Copyright © 2021 The Korean Society for Preventive Medicine
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Objectives
- Despite its advantages, it is not yet common practice in Korea for researchers to investigate disease associations using spatio-temporal analyses. In this study, we aimed to review health-related epidemiological research using spatio-temporal analyses and to observe methodological trends.
-
Methods
- Health-related studies that applied spatial or spatio-temporal methods were identified using 2 international databases (PubMed and Embase) and 4 Korean academic databases (KoreaMed, NDSL, DBpia, and RISS). Two reviewers extracted data to review the included studies. A search for relevant keywords yielded 5919 studies.
-
Results
- Of the studies that were initially found, 150 were ultimately included based on the eligibility criteria. In terms of the research topic, 5 categories with 11 subcategories were identified: chronic diseases (n=31, 20.7%), infectious diseases (n=27, 18.0%), health-related topics (including service utilization, equity, and behavior) (n=47, 31.3%), mental health (n=15, 10.0%), and cancer (n=7, 4.7%). Compared to the period between 2000 and 2010, more studies published between 2011 and 2020 were found to use 2 or more spatial analysis techniques (35.6% of included studies), and the number of studies on mapping increased 6-fold.
-
Conclusions
- Further spatio-temporal analysis-related studies with point data are needed to provide insights and evidence to support policy decision-making for the prevention and control of infectious and chronic diseases using advances in spatial techniques.
- The use of geographic information in the field of epidemiology was initiated by the renowned scientist John Snow, who linked a street water pump to the occurrence of cholera in Soho, London in the 1850s [1]. Since then, spatial analysis has grown to be an important part of epidemiology. However, its scope and applications were dramatically transformed by the advent of a new computing technology called geographic information systems (GIS) in the 1990s [2]. This technological innovation facilitated the application of spatial analyses and visualization techniques in the public health field [2,3]. GIS technology now allows us to visualize maps and assess spatial variability using more dynamic information [4], incorporating temporal information that changes over a short period [5,6]. These technological developments have encouraged many researchers to utilize more advanced spatio-temporal methodology, which takes into account both time and place [7].
- Prior research has reviewed the use of spatial analyses in the public health domain. In particular, Auchincloss et al. [8] reviewed studies that utilized spatial methods in epidemiology from 2000 to 2010. They chose 6 epidemiology journals and 1 public health journal for the review. Six types of spatial methods were included in their systematic review: proximity, aggregation, clustering, smoothing, interpolation, and spatial regression. Among these methods, proximity measures were most often used. A steep increase in the number of studies that used spatial methods was observed; however, these studies still constituted a small percentage (about 1%) of all published studies in public health.
- The use of spatial methods has increased in the scientific literature; however, little information has been published related to spatio-temporal analysis [8]. Spatio-temporal analysis has additional benefits over spatial analysis, as it considers both spatial and temporal correlations and makes it possible to observe the geographical and temporal distribution of phenomena through visualizations of epidemiological data that contain space and time information [9]. In addition, spatio-temporal modeling enables the investigation of trends over time, whereas spatial modeling only captures information at a specific point in time [10]. If the risk of diseases shows increasing or decreasing trends over time, then the use of spatio-temporal analysis is important. In addition, the appropriate use of space-time interactions can capture localized clusters that could be related to emerging environmental risks or consistent errors in the data recording process. This methodology also enables more immediate detection of persistent and uncommon patterns.
- Despite its advantages, it is not yet common practice in Korea for researchers to investigate disease relationships using spatiotemporal analyses, although the quantity of location-based data using mobile devices continues to increase, which has received substantial attention in public health research [1-3]. Moreover, to the best of our knowledge, while there have been some international reviews on the application of spatial analyses and spatio-temporal analyses in research, no such reviews have focused on research in Korea [2,6,8,11-13].
- Therefore, we conducted a comprehensive and systematic review on the application of spatial analyses, including GIS and spatio-temporal methods, in public health research in Korea. By reviewing the current state of the research and statistical methods, we will be able to evaluate the development of research methods and provide instant insights into potential opportunities in spatial and spatio-temporal research.
INTRODUCTION
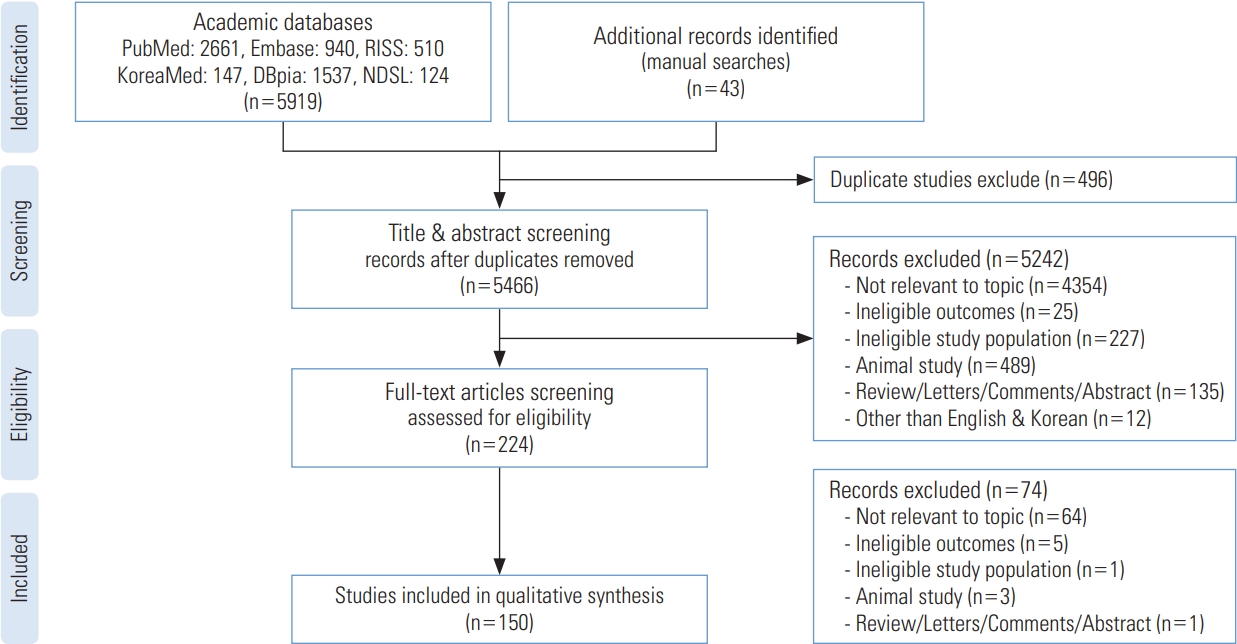
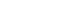
- The systematic review was conducted according to the PRISMA (Preferred Reporting Items for Systematic Review) checklist (Figure 1) [14].
- Search Strategy
- Health-related research that applied spatial or spatio-temporal methods were identified using 6 electronic databases: PubMed (https://pubmed.ncbi.nlm.nih.gov/) and Embase (https://www.embase.com/) as international databases, and KoreaMed (https://www.koreamed.org/), NDSL (http://www.ndsl.kr/), RISS (http://www.riss.kr), and DBpia (https://www.dbpia.co.kr/) as Korean academic databases. These databases were chosen for the following reasons: first, these databases are widely used by English-speaking or Korean-speaking researchers, and second, they allowed us to search using multiple keywords. Of note, some Korean databases did not allow searches with multiple keywords; hence we were limited to the 4 Korean search engines listed above. We searched for additional studies from Google Scholar and the reference lists of the final studies that we included in our review.
- Search terms included “disease mapping,” “spatial analysis,” “geographic information systems,” “spatio-temporal analysis,” “cluster analysis,” “epidemiology,” “disease,” “public health,” “Korea,” and term variants in English. Korean equivalents of those search terms were also used. Along with searching all the fields of the source documents from the databases, a hierarchically structured and controlled vocabulary thesaurus for biomedicine and life sciences—Medical Subject Headings for PubMed and Emtree for Embase—was used where applicable. The search was conducted using the advanced search option without any year limits. The full list of search terms is available in Supplemental Material 1.
- Inclusion and Exclusion Criteria
- Our eligibility criteria were that the studies had to be original, published studies written in Korean or English focusing on the 3 major areas relevant for this systematic review. These areas were (1) the use of specific analytical techniques—in our case, the use of GIS, spatial analysis, or spatio-temporal analysis; (2) a focus on the field of public health; and (3) the use of Korean population data and not a focus on the global population. Studies were included in the review if they met the following criteria: (1) human-subject research or human-related disease; (2) a main focus on public health (i.e., studies that only discussed methods of spatial or spatio-temporal analysis were excluded); (3) health-related outcomes; and (4) Korean population data (Koreans residing in Korea). The search was limited to studies that were written in English or Korean. Studies that did not meet these criteria were excluded from the review.
- Data Collection and Processing
- We downloaded all the available data using Endnote X9 (Clarivate Analytics, Philadelphia, PA, USA). Two reviewers (HGB and NL) independently identified eligible studies by reviewing the titles and abstracts, and we then assessed the full text of the studies that needed more information for eligibility identification. All the studies were cross-checked by both researchers. All studies with unresolved eligibility identification were reviewed by the third reviewer (SSH), and a consensus among all researchers was reached.
- Reviewing Selected Studies
- We extracted the following data from each study that was included: (1) study information (first author’s name, first author’s affiliation, the title of a study, year of publication, and language); (2) publication information (name of the published journal, each journal’s research area classification, and whether the journal was domestic or international); (3) study characteristics (study design, subject of research, research goal, data source, and data information); and (4) methods (geographical or temporal unit of analysis, the software used for analysis, types of spatial methods, and the presence of regression analysis). Due to the large number of included studies and extracted data, important information regarding the included studies is organized and presented in Supplemental Materials 2 and 3.
- Ethics Statement
- This paper is a systematic review; therefore, it did not require ethical approval.
METHODS
- The flow chart of the study selection process is shown in Figure 1. The total number of collected studies from the databases was 5919. Of these, 3601 studies were from international databases (PubMed: 2661; Embase: 940) and 2318 from Korean databases (KoreaMed: 147; DBpia: 1537; RISS: 510; NDSL: 124). Forty-three additional studies were found from manual searches. After the removal of duplicates (n= 496), we excluded studies with non-relevant topics (n=4354), studies with ineligible outcomes (n=25), studies with ineligible populations (n=227), non-human studies (n=489), studies that were not original research (n=135), and non-English or Korean studies (n=12) by screening titles and abstracts. Next, we screened the full-text of studies to assess whether they met our eligibility criteria, and through this process, we excluded 74 studies. Ultimately, 150 studies were included in the final list for the systematic review.
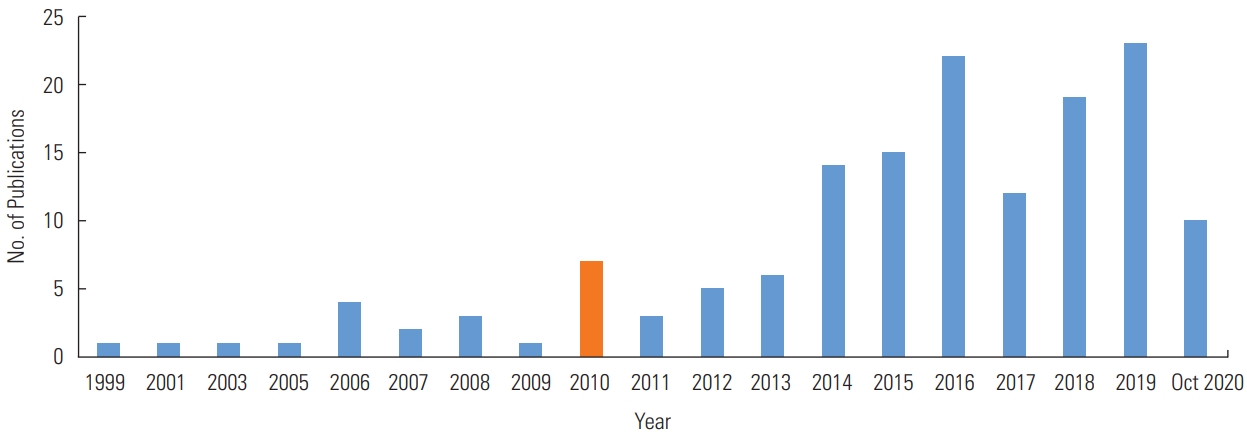
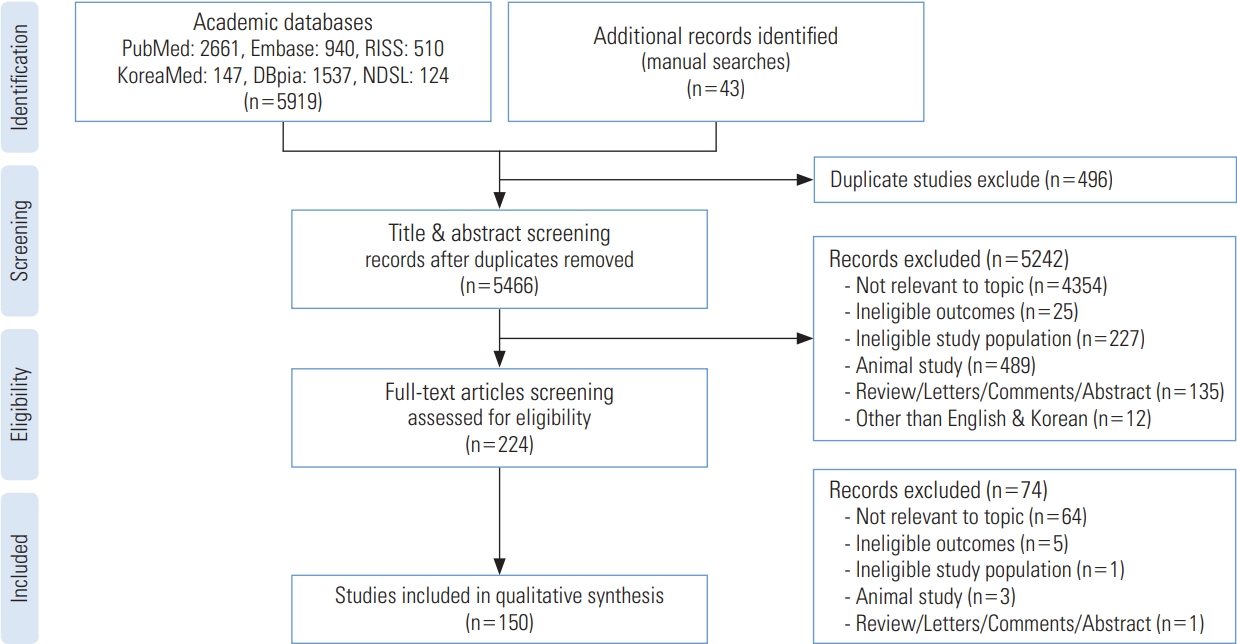
- Figures 2 to 4 illustrate the characteristics of the studies included in the final list. From 1999 to the present, the number of publications increased steadily, as shown in Figure 2. We analyzed the studies using 2010 as a benchmark because the number of publications started to rise in 2010 and peaked in 2019 (n=23), although ongoing research is expected to be published in the future. The number of studies published after 2010 (n=129) was over 7-fold higher than the number of studies published before 2010 (n=21).
- Study Designs and Exposures
- More than half (n=95; 63.3%) of the studies were ecological studies, while 32.0% (n=48) were cross-sectional studies. The ecological studies used exposures or outcomes measured at the group level. Conversely, cross-sectional studies measured exposure and disease outcomes in a single study at the individual level. Among the 95 studies using ecological designs, the exposure was related to the environment in 21 and health behavior in 1. Of the cross-sectional studies, 25.0% (n=12) were related to environmental exposures, and 1 study dealt with health behavior. Few studies (n=3) used a cohort study design, and 1 study used each of the following types of study design: intervention study, nested case-control study, retrospective observational study, and retrospective cohort study.
- The most common field of the journals where the articles were published was public health, with 42.0% (n=63), followed by 20.0% (n=30) for medical journals, including preventive or emergency medicine, and 8.0% (n=12) for geography. Domestic academic journals accounted for 89 studies, which exceeded the number of articles published in international academic journals. Since 2010, domestic academic journals published 17 to 72 studies, and international journals published 4 to 57 studies, indicating that research using spatial analysis was actively carried out in the public health field.
- Study Topic and Data Type
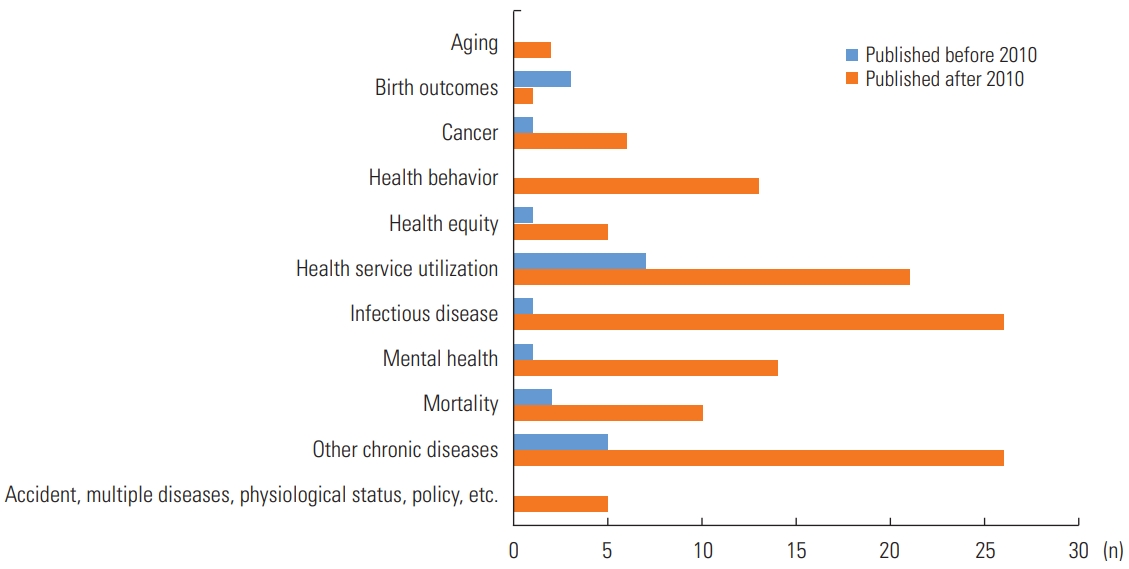
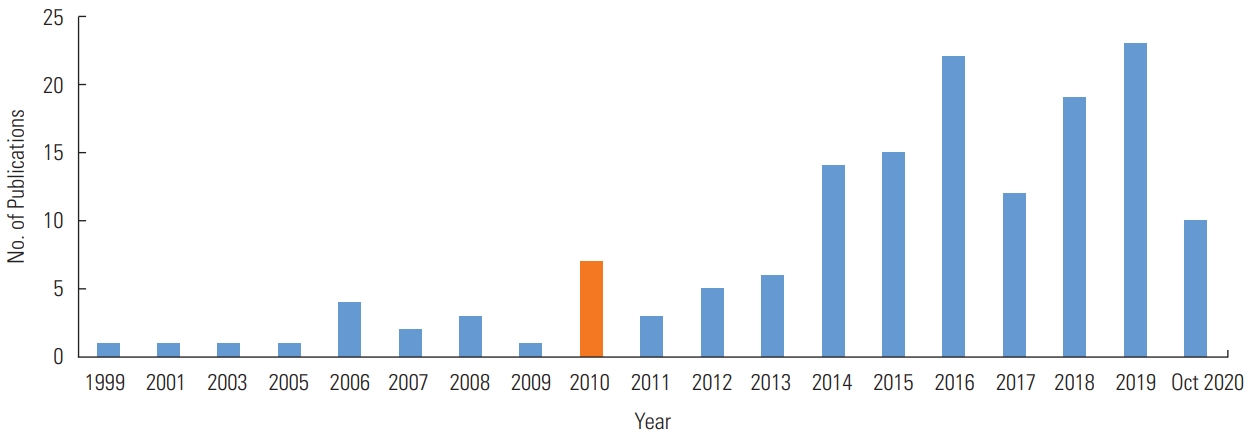
- Figure 3 demonstrates a further classification by research topic. Most of the studies before 2010 focused on health service utilization (n=7) and chronic diseases (n=5); however, after 2010, chronic diseases (n=26), infectious disease (n=26), and health service utilization (n=21) comprised 56.6% of the studies. In addition, as the major issues changed with time, spatial analysis was conducted on new research topics after 2010. A noteworthy research topic was aging (n=2), as well as other topics, including accidents, multiple diseases, or psychological status (n=5). A noticeable increase was found in studies on infectious diseases, mental health, and health-related topics (e.g., service utilization, equity, or behavior) from before 2010 to after 2010. Among the 150 included studies, 38 studies (25.3%) focused on environmental exposures, and 2 studies were related to health behaviors such as smoking and suicide.
- Most studies used areal data compiled as aggregate regions (n=129, 86.0%), whereas very few studies used point data (n=21, 14.0%). Of the studies using point data, 3 (14.3%) used spatial methods implying clustering only, and 52.4% (n=11) used the proximity method, either in isolation or with other techniques. The geographic units of analyses were provinces (n=10, 6.7%), counties (n=75, 50.0%), and towns (n=44, 29.3%). However, 14.0% of the included studies were classified as “unknown” in this regard because they did not describe the geographic units used.
- Types of Spatial Methods and Software
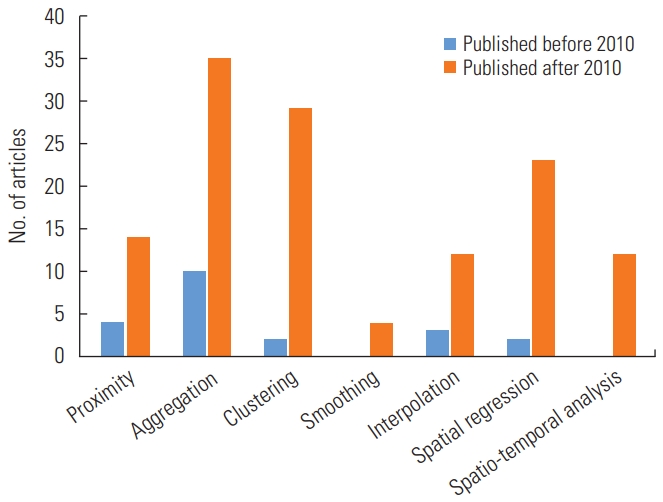
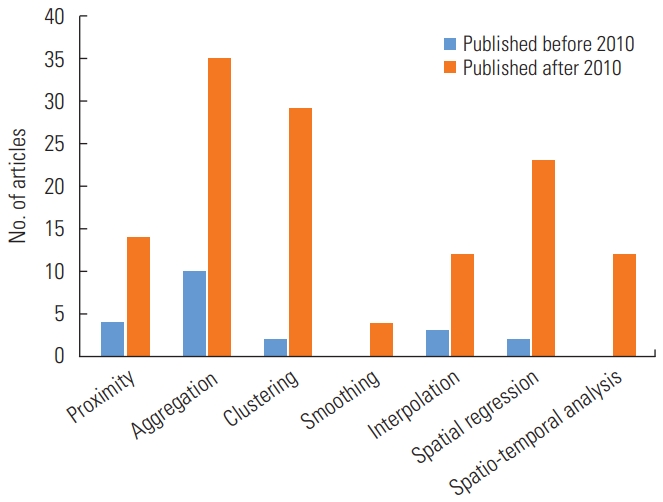
- In the analysis of types of spatial methods and software, we allowed duplicate counts to reflect studies that used multiple different spatial methods or software programs. As shown in Figure 4, 10 studies used aggregation before 2010, whereas after 2010, the majority of studies used aggregation (n=35), spatial clustering (n=29), and spatial regression (n=23) as the methods. Furthermore, no studies used spatio-temporal analyses before 2010, and relatively few studies applied spatio-temporal analyses after 2010 (n=15). The total number of studies that only used 1 spatial method was 88 (58.7%), while 35 studies (23.3%) used 2 methods, 25 studies (16.7%) used 3 methods, and only 1 study utilized four methods. In terms of visualization, approximately 78.7% (n=118) of the included studies published after 2010 presented disease mapping, providing useful information in detail.
- ArcGIS (n=49, 32.7%) was the most frequently used GIS software. Sixteen studies (10.7%) used GeoDa, 5 (3.3%) used QGIS, 11 (7.3%) used ArcMap, 5 (3.3%) used ArcView, 6 (4.0%) used SatScan, and 2 (1.0%) used MapWizard. Of the 150 studies, 30 studies used R, 28 studies used SAS, 22 studies used SPSS, and 11 studies used WinBUGS. Many studies used more than 1 software program for analysis (n=105, 70.0%). Some studies used other software such as WindNinja and the Joinpoint Regression Program. More details are provided in Supplemental Material 3.
RESULTS
- With advances in technology and methodology, research using space-time analysis methods will continue to expand. Through the application of these methods in various research fields, data use and analysis using programs have become easier. This systematic review of studies related to health and disease outcomes in Korea from a public health point of view using spatial or spatio-temporal analysis analyzed 150 studies. There was an explosive increase in spatial research after 2010. ArcGIS was the most commonly used program, followed by R, SAS, and SPSS. The most frequently used methods were aggregation, spatial clustering, and spatial regression, while smoothing and interpolation were used less frequently.
- Under-utilization of Surveillance Data
- According to the Centers for Disease Control and Prevention, surveillance data allow health-related public health surveillance by estimating the scale of a particular problem, determining the distribution of diseases, generating hypotheses, and monitoring changes [15]. Surveillance data are often used for early warnings and health monitoring [16], and conducting spatio-temporal analyses of surveillance data is at the core of preventive medicine [17]. Such analyses can also help reduce bias and uncertainty regarding spatio-temporal variation related to health outcomes [18]. As an example of surveillance data, the Korea Disease Control and Prevention Agency operates a thermal disease emergency room monitoring system to mitigate the health impacts of heat waves and an infectious disease monitoring system to manage and prevent infectious diseases such as cholera or dengue fever [19]. Nonetheless, the utilization of surveillance data to implement spatial analysis was quite low, as only 4 studies used surveillance data. Most of the studies relied heavily on cross-sectional surveys such as the Korea National Health and Nutrition Examination Surveys and Korea Community Health Survey, as well as Korean Statistical Information Service data; these are regularly conducted, well-known, and easily-found surveys that contain large-scale data representing the Korean civilian population and enable comparisons of difference between provinces. Nevertheless, analysis using surveillance data provides a better understanding of risk factors and is more suitable for comparisons of the epidemiological patterns of infectious diseases considering periods and patterns [20]. Utilizing emergency room-based data using SatScan can enable the expansion of the public health monitoring system. Emergency medicine and infectious diseases are fields that could benefit greatly from spatial and spatio-temporal epidemiology. Because space-time disease surveillance systems can serve as a superb tool for improving public health [16], using surveillance data in the field of spatial and spatiotemporal epidemiology will be advantageous.
- Lack of Small-area Analyses
- Using appropriate statistical methodology, high-resolution geographically referenced data can facilitate investigations of local geographic variations in diseases [4]. As an example of geo-reference data using big data and GIS in healthcare and public health, thematic maps and interactive statistical maps are provided through the Statistical Geographic Information Service and the Korean National Health Insurance Service-Atlas, and the Emergency Medical Service System operated by the National Emergency Medical Center provides health maps and provides visualizations of access to local medical resources. These kinds of datasets offer opportunities for small-area, town-level analyses; however, we still rely heavily on county-level analyses. A major challenge in small-area analysis is the preservation of participants’ privacy and confidentiality [4]. Furthermore, we found that only a few studies used point data, whereas most used areal data. In light of limitations in the availability of data that researchers can use, it is difficult to identify spatial barriers or boundaries reflecting difference boundaries [21], leading to the modifiable areal unit problem (MAUP), which makes it challenging for research to fully reflect the spatial characteristics of the phenomenon under investigation. The MAUP refers to the fact that aggregating data into different sizes or geographical units for spatial analysis can cause many problems, including accuracy, scale, quality (possible bias) and confounding factors [22]. One of the best methods to solve the MAUP is to use data that can provide details on spatial units at the small-area level. When checking cluster patterns, it is difficult to guarantee the reliability of the study results unless the MAUP is dealt with [23].
- Open-source Software
- Rapid growth in tools for spatial data analysis has occurred since the 1980s [24]. Through these remarkable developments, made possible by the efforts of numerous researchers, software that users can use for free has also been developed. R and QGIS are examples of free, publicly accessible open-source software (OSS). OSS is distributed to anyone to use, run, modify, and redistribute freely, thereby saving research costs, which is especially important in developing countries [5]. With OSS and studio programs, many researchers can apply diverse research topics with new perspectives. In addition, researchers can share their open-source code and packages without licensing fees, thereby facilitating the process, analysis, and presentation of research. Web-based spatial information systems can also be created using OSS; compared with commercial software, they are simple and compatible with other operating systems [6]. Despite the development of OSS, we found that ArcGIS was the most frequently used GIS tool in the reviewed studies. If researchers who conduct spatial analysis share open-source code, it will increase the frequency of usage, making it possible to build infrastructure related to spatial data in the future [25] to promote multidisciplinary research.
- Policy Implications
- The number of research studies using spatio-temporal analysis has increased since 2010, especially regarding health service utilization, infectious disease, and other chronic diseases. Nevertheless, there are still no epidemiological studies that have used a spatio-temporal analysis as a basis for policy development. Because spatio-temporal analysis incorporates spatial and temporal corrections to determine the cause or clustering of a disease, there is a need to strengthen this research area to develop policies to prevent community-based chronic diseases and acute and non-acute infectious diseases. An institutional approach is needed to protect and strengthen the privacy protections of personal information that can be accessed during this process. Currently, the Personal Information Protection Commission collects data on personal information problems caused by epidemiological investigations during the coronavirus disease 2019 (COVID-19) pandemic, and the Infectious Disease Control Act was revised in March 2021 to consider privacy concerns.
- Since the start of the COVID-19 pandemic, 63 scientific articles have been published with geospatial and spatial-statistics analyses [26]. A review of the latest literature revealed that understanding spatio-temporal dynamics, as discussed herein, can help identify the scope and impact of infectious diseases, contributing to efforts to reduce disease spread. From a multidisciplinary perspective, research should more actively conduct spatial and spatio-temporal analyses in epidemiology to prepare for global pandemics and infectious diseases.
DISCUSSION
- Relatively few studies have been published on applications of spatial and temporal analyses related to public health issues in Korea. Moreover, there is a lack of data from small-area unit analyses in the field of spatio-temporal epidemiology. Further studies related to spatio-temporal analysis with point data are needed to provide insights and evidence to support policy decision-making for the prevention and control of infectious and chronic diseases using advances in spatial techniques. Further follow-up research is also recommended considering the increasing use of geocoding.
CONCLUSION
SUPPLEMENTAL MATERIALS
Supplemental Material 2
Supplemental Material 3
-
CONFLICT OF INTEREST
The authors have no conflicts of interest associated with the material presented in this paper.
-
FUNDING
This research was funded by Seoul National University’s New Faculty Research Resettlement Fund (Grant No: 900-20170068) and supported by a National Research Foundation of Korea grant funded by the Korean government (No.21B20151213037).
Notes
ACKNOWLEDGEMENTS
-
AUTHOR CONTRIBUTIONS
Conceptualization: HGB, SSH. Data curation: HGB, NL. Funding acquisition: SSH. Methodology: HGB, NL, SSH. Writing – original draft: HGB. Writing – review & editing: HGB, NL, SSH.
Notes
- 1. Snow J. On the mode of communication of cholera. Edinb Med J 1856;1(7):668-670PubMed
- 2. Meliker JR, Sloan CD. Spatio-temporal epidemiology: principles and opportunities. Spat Spatiotemporal Epidemiol 2011;2(1):1-9ArticlePubMed
- 3. Pickle LW. A history and critique of U.S. mortality atlases. Spat Spatiotemporal Epidemiol 2009;1(1):3-17ArticlePubMed
- 4. Castronovo DA, Chui KK, Naumova EN. Dynamic maps: a visual-analytic methodology for exploring spatio-temporal disease patterns. Environ Health 2009;8: 61ArticlePubMedPMC
- 5. Nadi S, Delavar MR. Spatio-temporal modeling of dynamic phenomena in GIS. In: ScanGIS 2003: Proceedings 9th Scandinavian Research Conference on Geographical Information Sciences. 2003 Jun 4-6; Espoo, Finland: 2013, p. 215-225
- 6. Pfeiffer DU, Stevens KB. Spatial and temporal epidemiological analysis in the Big Data era. Prev Vet Med 2015;122(1-2):213-220ArticlePubMedPMC
- 7. Musa GJ, Chiang PH, Sylk T, Bavley R, Keating W, Lakew B, et al. Use of GIS mapping as a public health tool-from cholera to cancer. Health Serv Insights 2013;6: 111-116ArticlePubMedPMC
- 8. Auchincloss AH, Gebreab SY, Mair C, Diez Roux AV. A review of spatial methods in epidemiology, 2000-2010. Annu Rev Public Health 2012;33: 107-122ArticlePubMedPMC
- 9. Yu G, Yang R, Wei Y, Yu D, Zhai W, Cai J, et al. Spatial, temporal, and spatiotemporal analysis of mumps in Guangxi Province, China, 2005-2016. BMC Infect Dis 2018;18(1):360ArticlePubMedPMC
- 10. Richardson S, Abellan JJ, Best N. Bayesian spatio-temporal analysis of joint patterns of male and female lung cancer risks in Yorkshire (UK). Stat Methods Med Res 2006;15(4):385-407ArticlePubMed
- 11. Dijkstra A, Hak E, Janssen F. A systematic review of the application of spatial analysis in pharmacoepidemiologic research. Ann Epidemiol 2013;23(8):504-514ArticlePubMed
- 12. Robertson C, Nelson TA, MacNab YC, Lawson AB. Review of methods for space-time disease surveillance. Spat Spatiotemporal Epidemiol 2010;1(2-3):105-116ArticlePubMedPMC
- 13. Roddick JF, Hornsby K, Spiliopoulou M. An updated bibliography of temporal, spatial, and spatio-temporal data mining research. In: International Workshop on Temporal, Spatial, and Spatio-Temporal Data Mining. Heidelberg: Springer; 2000. p. 147-163
- 14. Vrabel M. Preferred reporting items for systematic reviews and meta-analyses. Oncol Nurs Forum 2015;42(5):552-554ArticlePubMed
- 15. Institute of Medicine (U.S.) Committee on a National Surveillance System for Cardiovascular and Select Chronic Diseases. A nationwide framework for surveillance of cardiovascular and chronic lung diseases. Washington, DC: National Academies Press; 2011. p. 65-90
- 16. Hornsby K, Egenhofer MJ. Identity-based change: a foundation for spatio-temporal knowledge representation. Int J Geogr Inf Sci 2000;14(3):207-224Article
- 17. Ward MP. Spatio-temporal analysis of infectious disease outbreaks in veterinary medicine: clusters, hotspots and foci. Vet Ital 2007;43(3):559-570PubMed
- 18. Blangiardo M, Boulieri A, Diggle P, Piel FB, Shaddick G, Elliott P. Advances in spatiotemporal models for non-communicable disease surveillance. Int J Epidemiol 2020;49(Suppl 1):i26-i37ArticlePubMedPMC
- 19. Park S, Cho E. National infectious diseases surveillance data of South Korea. Epidemiol Health 2014;36: e2014030ArticlePubMedPMC
- 20. Pfeiffer DU, Minh PQ, Martin V, Epprecht M, Otte MJ. An analysis of the spatial and temporal patterns of highly pathogenic avian influenza occurrence in Vietnam using national surveillance data. Vet J 2007;174(2):302-309ArticlePubMed
- 21. Li P, Banerjee S, Hanson TA, McBean AM. Bayesian models for detecting difference boundaries in areal data. Stat Sin 2015;25(1):385-402PubMedPMC
- 22. Souris M. Epidemiology and geography : principles, methods and tools of spatial analysis. Hoboken: John Wiley & Sons; 2019. p. 221
- 23. Choi DJ, Suh YC. A study on the exploratory spatial data analysis of the distribution of longevity population and the scale effect of the modifiable areal unit problem (MAUP). J Korean Assoc Geogr Inf Stud 2013;16(3):40-53. (Korean)Article
- 24. Anselin L, Syabri I, Kho Y. GeoDa: an introduction to spatial data analysis. In: Fischer M, Getis A, edtitors. Handbook of applied spatial analysis. Berlin: Springer; 2010. p. 73-89
- 25. Hall GB, Leathy MG. Open source approaches in spatial data handling. Berlin: Springer; 2008. p. 1-20
- 26. Franch-Pardo I, Napoletano BM, Rosete-Verges F, Billa L. Spatial analysis and GIS in the study of COVID-19. A review. Sci Total Environ 2020;739: 140033ArticlePubMedPMC
REFERENCES
Figure & Data
References
Citations

- Group I pharmaceuticals of IARC and associated cancer risks: systematic review and meta-analysis
Woojin Lim, Sungji Moon, Na Rae Lee, Ho Gyun Shin, Su-Yeon Yu, Jung Eun Lee, Inah Kim, Kwang-Pil Ko, Sue K. Park
Scientific Reports.2024;[Epub] CrossRef - Spatiotemporal models of dengue epidemiology in the Philippines: Integrating remote sensing and interpretable machine learning
Don Enrico Buebos-Esteve, Nikki Heherson A. Dagamac
Acta Tropica.2024; 255: 107225. CrossRef - Spatio-Temporal Analysis of Leptospirosis Hotspot Areas and Its Association With Hydroclimatic Factors in Selangor, Malaysia: Protocol for an Ecological Cross-sectional Study
Muhammad Akram Ab Kadir, Rosliza Abdul Manaf, Siti Aisah Mokhtar, Luthffi Idzhar Ismail
JMIR Research Protocols.2023; 12: e43712. CrossRef - Epidemiological characteristics and spatiotemporal analysis of mumps at township level in Wuhan, China, 2005–2019
Ying Peng, Peng Wang, De-guang Kong, Wen-zhen Li, Dong-ming Wang, Li Cai, Sha Lu, Bin Yu, Bang-hua Chen, Pu-Lin Liu
Epidemiology and Infection.2023;[Epub] CrossRef - A Systematic Review of Areal Units and Adjacency Used in Bayesian Spatial and Spatio-Temporal Conditional Autoregressive Models in Health Research
Zemenu Tadesse Tessema, Getayeneh Antehunegn Tesema, Susannah Ahern, Arul Earnest
International Journal of Environmental Research and Public Health.2023; 20(13): 6277. CrossRef - Use of geographically weighted regression models to inform retail endgame strategies in South Korea: application to cigarette and ENDS prevalence
Heewon Kang, Eunsil Cheon, Jaeyoung Ha, Sung-il Cho
Tobacco Control.2023; : tc-2023-058117. CrossRef - EpiVECS: exploring spatiotemporal epidemiological data using cluster embedding and interactive visualization
Lee Mason, Blànaid Hicks, Jonas S. Almeida
Scientific Reports.2023;[Epub] CrossRef - Spatiotemporal Trends and Distributions of Malaria Incidence in the Northwest Ethiopia
Teshager Zerihun Nigussie, Temesgen T. Zewotir, Essey Kebede Muluneh, Wei Wang
Journal of Tropical Medicine.2022; 2022: 1. CrossRef - Small-scale spatiotemporal epidemiology of notifiable infectious diseases in China: a systematic review
Junyao Zheng, Guoquan Shen, Siqi Hu, Xinxin Han, Siyu Zhu, Jinlin Liu, Rongxin He, Ning Zhang, Chih-Wei Hsieh, Hao Xue, Bo Zhang, Yue Shen, Ying Mao, Bin Zhu
BMC Infectious Diseases.2022;[Epub] CrossRef - Characteristics of Atmospheric Compounds based on Regional Multicorrelation Analysis in Honam Area
Sung-Hyun Oh, Sea-Ho Oh, Min-Suk Bae
Journal of Environmental Analysis, Health and Toxicology.2022; 25(3): 85. CrossRef - Spatiotemporal analyses of the epidemiological characteristics of diabetes mellitus
Sang Youl Rhee
Epidemiology and Health.2021; 43: e2021102. CrossRef
 KSPM
KSPM


 PubReader
PubReader ePub Link
ePub Link Cite
Cite